Each time a protein is produced by the ribosome, errors are happening at a detectable rate (Mohler & Ibba, 2017). One type of error is ribosomal frameshifting, where the ribosome ‘slips’ on the mRNA into a different reading frame. As a result, a completely different peptide sequence is produced (Caliskan et al., 2015). Detecting frameshifted sequences is challenging for two reasons:
1) Frameshifting generally happens rarely, and the resulting non-canonical proteins will be of very low abundance.
2) The frameshifted proteins differ a lot from the annotated sequences, thus they will not be detected when matched against a normal protein database.
For these reasons, even in model organisms, there is no proteome-wide picture of the extent of frameshifting. We want to find out how many proteins are affected by this phenomenon, and at what rates frameshifts are happening. We use S. cerevisiae as a simple eukaryotic model.
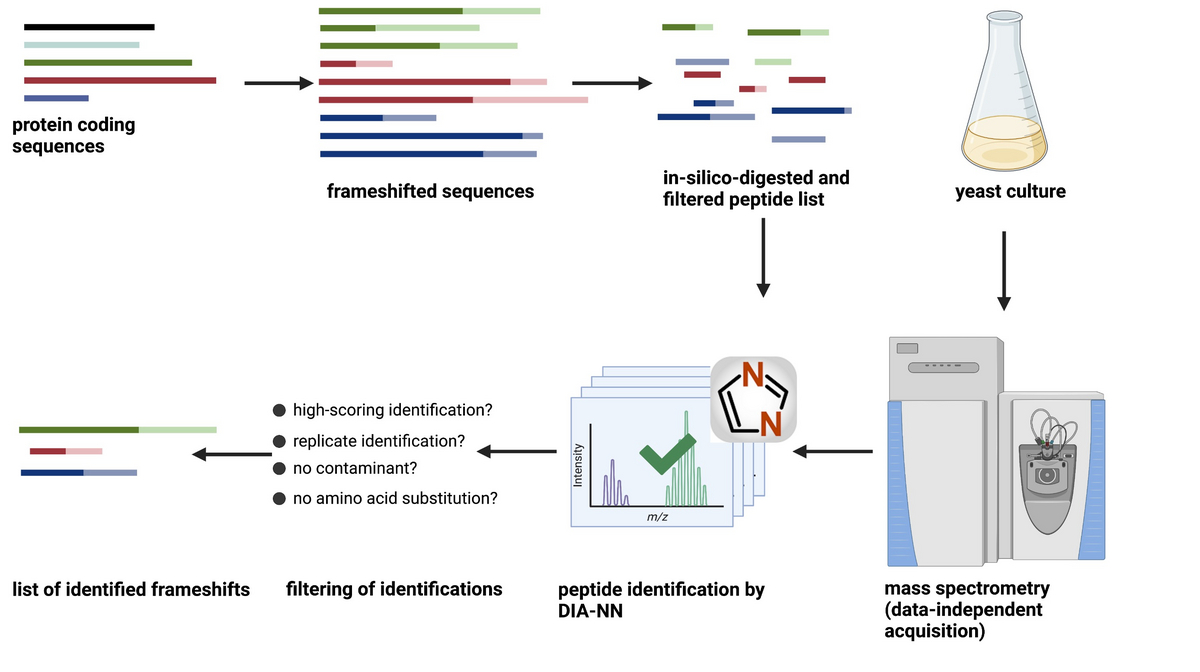
To identify novel frameshifts, we first predict, based on the coding sequences, the protein sequences that would result from a frameshift at any position. This very large sequence database is digested in-silico, filtered and annotated such that every peptide identifies a unique frameshifting event. To screen for all possible frameshift peptides, we apply a data-independent acquisition (DIA) scheme with very narrow, overlapping isolation windows. This way, we achieve high specificity without sacrificing proteome coverage.
After the proteome-wide screen, candidate frameshift peptides are validated and quantified (relative to the canonical protein) with the help of isotopically labelled, chimeric proteins containing the target peptides (FastCAT, Rzagalinski et al., 2022). We were able to identify and quantify several novel frameshift events in yeast, with frameshift rates ranging from 1 to 10%.
Agnes Toth-Pectroczy, Cedric Landerer (Toth-Petroczy lab, MPI CBG)
Aliona Bogdanova (PEPC, MPI CBG)

Contact: poehls@mpi-cbg.de