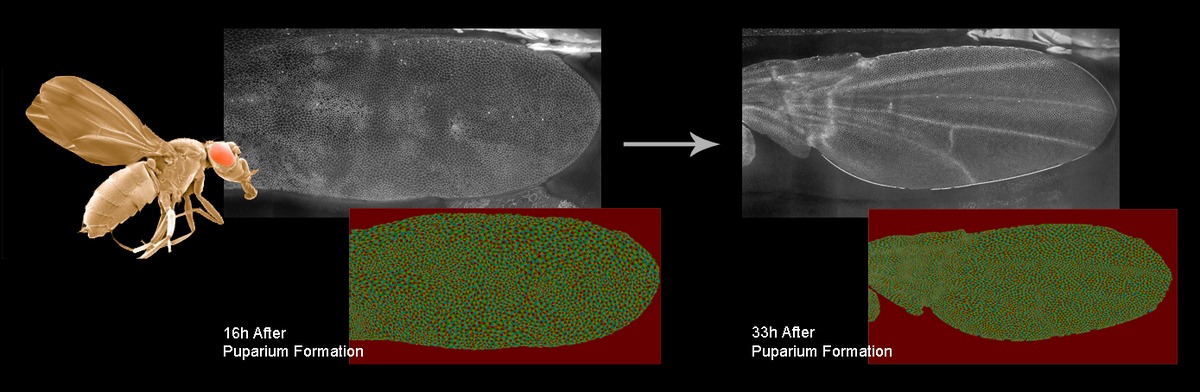
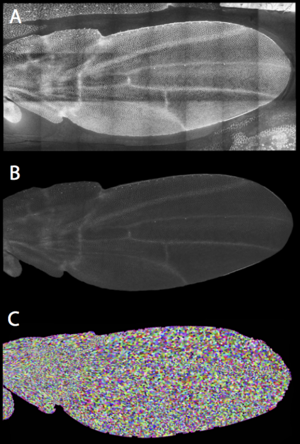
Organ development is characterized by a high degree of proliferation and spatial rearrangement of its constituent cells. For example, during wing development in Drosophila, cells undergo oriented elongation, division and rearrangements to generate the final shape of the wing. For a detailed understanding of this process, one need to measure the shape, number and cell polarity of all cells over time. Although the visualization of this complete developmental process is possible using fluorescent light microscopy, it is not yet possible to extract these quantitative properties with an efficiency that allows for hundreds or thousands of recordings, as might be needed in a perturbation study.
The goal of this project is the development of highly accurate yet efficient algorithms to address this problem. An automated method will be provided to detect every cell boundary (segmentation) and to follow each cell through time (tracking).