Endocytosis is an essential process serving multiple key cellular functions, such as nutrient uptake, signal transduction, and defense against pathogens. In endocytosis, a plasma membrane patch is internalized forming a transport vesicle containing cargo from the extracellular space. Transport vesicles bud off from the plasma membrane and eventually fuse with a target cellular compartment. To ensure correct cargo delivery, transport vesicles must be highly selective in recognizing the correct target membrane for fusion. Therefore, the vesicles harbor specific surface markers identifying their origin and type of cargo, whereas target membranes display the corresponding receptors. In this process, Rab proteins direct the vesicle to the specific spot on the target membrane, whereas SNARE proteins mediate membrane fusion. Before both membranes can fuse, they are recognized and pulled close together by proteins called membrane tethers.
Currently, the molecular mechanisms underlying the transition from membrane tethering to fusion remain elusive. Using an in vitro reconstitution assay in vesicles, our lab found that the Rab5 machinery is required for endosome membrane fusion, for which SNAREs alone are insufficient. Interestingly, the Rab5 machinery couples membrane recognition to mechanics. Rab5 binding to the tethering factor EEA1 induces an allosteric conformational change, generating an entropic collapse force that we measured with the help of optical tweezers. This force might help pulling the membranes together before they finally fuse.
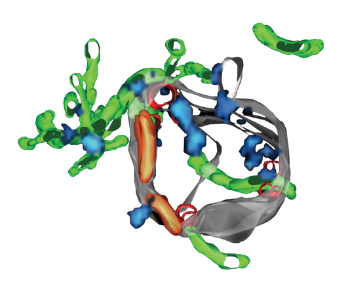
Complementary to this approach, we studied the structural organization of the Rab5 machinery on early endosomes. By correlative super-resolution and electron microscopy (SuperCLEM), we found that Rab5 and its effectors form distinct domains on the endosomal membrane. We have now succeeded in reconstituting such domains in vitro with a minimal machinery including Rab5, Rab GDI and the Rabaptin5-Rabex5 complex. The combination of in vitro and in vivo systems allows us to answer questions regarding the formation, dynamics and role of Rab5 domains in the context of endosome structure and function.
No cell is an island – and therefore we must understand molecular processes within the context of a tissue. In our group, we study the liver because its remarkable tissue features are perfectly fit to answer our research questions.
A large part of the liver consists of parenchymal cells called hepatocytes that are honeycombed by two 3D microcirculatory systems: the sinusoids that transport blood and the bile canaliculi network that drains bile into bile ducts. Hepatocytes are polarized cells, meaning that there are functional and spatial differences within the cell. Simple polarized cells are quite common and form epithelia, such as the gut, with a single apical surface facing the lumen and a basal surface facing the blood. However, hepatocytes possess a belt-like apical membrane that wraps around the entire cell. The apical surfaces of several hepatocytes pair with the neighboring ones to form a small lumen in between. In its entirety, these connected lumina form the bile canalicular network. In our group, we study the molecular and physical mechanisms underlying hepatocyte polarization and, thus, bile canaliculi formation.
We recently identified small apical membranous structures in hepatocytes that we termed “apical bulkheads”. These apical membrane extensions project inside the lumen of bile canaliculi and we believe that they might contribute to bile canaliculi formation and elongation. Interestingly, we found the small GTPase Rab35, usually known to be involved in membrane trafficking, to be involved in this process. Upon Rab35 knockdown, apical bulkheads were lost and instead of elongated bile canaliculi, hepatocytes formed spherical cysts and tubes similar to epithelial cells. Therefore, Rab35 plays an unexpected new biomechanical function in hepatocyte polarity and the formation of the bile canaliculi network. Currently, we are investigating the role of Rab35 and apical bulkheads in cell polarity and in tissue homeostasis.
The liver is the largest metabolic organ of the human body and vital for detoxification. Although the general organization of the liver in millimeter-sized lobules is well understood, the tissue organization within the lobule remains enigmatic. It was long believed that sinusoidal endothelial cells and hepatocytes form a heterogeneous parenchyma intersected by labyrinths of sinusoids and bile canaliuli without apparent order. However, for the liver to work properly, each hepatocyte needs to be in contact with both networks, so that the two networks can never intersect. This poses the question of self-organization to satisfy these competing design requirements. Hans Elias pioneered this model of the liver lobule structure in 1949, but since then very little progress had been made.
In our group, we use computer-aided image analysis techniques to understand how liver cells assemble to build tissue from a structural perspective. We developed an image analysis pipeline to process high-resolution microscopy images of liver samples achieving a comprehensive and accurate 3D digital representation of the tissue. Based on these findings, we developed a predictive 3D multi-scale model that simulates bile fluid dynamics and regulation of organ size during regeneration.
With this system, we defined new organizing principles of the liver lobule that go beyond the structural model postulated by Hans Elias in 1949. In a highly collaborative work, we showed that hepatocytes are partially oriented along a common axis. The orientation of individual cells can differ, however the total of cells exhibits a certain order. This concept is used to describe e.g. liquid crystal displays (LCDs) in engineering. We found that hepatocytes in liver lobules organize following a long-range 3D liquid crystal order.
With two distinct approaches, we are currently dissecting the molecular mechanisms whereby liver cells interact and self-organize. First, we are reconstituting a versatile cellular in vitro system that we can use to manipulate candidate factors in tissue organization selectively. In a second approach, we are focusing on tissue repair and remodelling during liver regeneration. In collaboration with the research group of Meritxell Huch, we successfully generated liver-derived organoids which are promising tools in this endeavour.
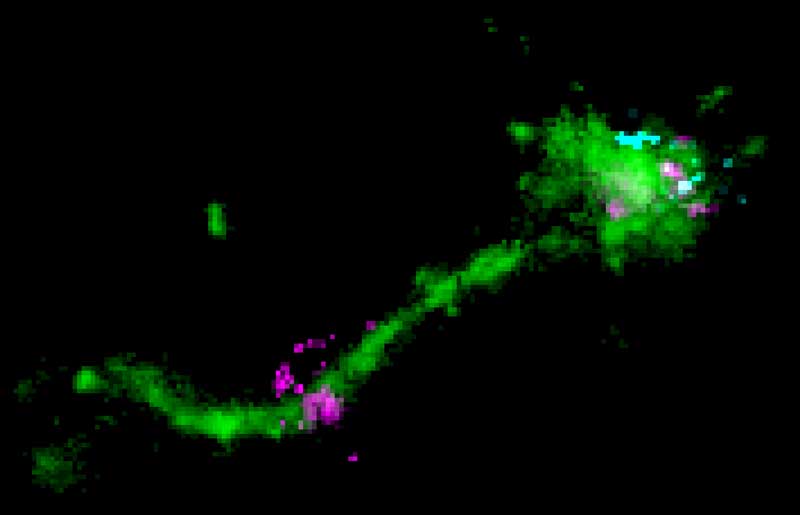
Macromolecules such as nucleic acids or proteins possess a great potential as therapeutics in medical applications and research. However, their entry in the cell and targeted delivery to their specific sites of action is still very problematic and mechanisms are unclear. Cells take up and traffic macromolecules via the endocytic pathway. Currently, uptake and site-specific targeting are major bottlenecks in macromolecule delivery. The cytoplasm is especially difficult to target, since the macromolecule needs to escape the endosome along the trafficking route.
Over the last years, our lab has been exploring uptake mechanisms and endosomal escape in various collaborations with pharmaceutical companies. We have characterized the uptake and endosomal distribution of siRNAs and identified small molecules that enhance the efficacy of delivery systems provided by our collaborators. Our expertise in endosomal trafficking and membrane fusion allows us to use bio-inspired approaches to improve macromolecule delivery systems.