Alex Kalinka, Melek Asli Kayserili, Christopher Schmied, Vineeth Surendranath
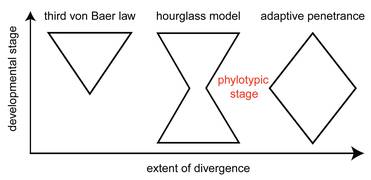
We are interested in quantitatively testing the "hourglass model" of morphological evolution, which postulates that divergent animal species that belong to one animal phylum pass through a similar period of development known as the "phylotypic period." The hourglass model builds on classic observations of von Baer, Darwin and Haeckel on morphological embryonic similarity among animal species and recognizes that early stages of development before the phytotypic perid are often accomplished by surprisingly variable mechanisms. Although this long-standing model describes a fundamental pattern in the evolution of animal development, the validity of the model and its mechanistic underpinning remains in contention.
We collected micro-array time-courses for embryogenesis of six species of fruitflies using custom designed species-specific Agilent arrays interrogating approximately 3500 genes expressed during embryogenesis. Statistical and quantitative genetic analysis of the data indicates that gene expression levels vary among species more in the beginning and at the end of embryogenesis compared to mid-embryogenesis, in agreement with the predictions of the hourglass model (Kalinka, Varga et al. 2010).
We synthesized data from scientific literature on divergence of early developmental processes and the underlying gene regulatory networks with recent comparative genomics studies in various phyla and conclude that indeed there is ample evidence that early development is evolutionarily labile. We seek explanation for the early divergence in ecological circumstances of the parental generation (especially mothers who invest maternal resources into the early embryos) that result in selection for divergent life history strategies (Kalinka and Tomancak 2012).
In the future, we plan to expand the comparative inter-species microarray analysis to the whole genome using RNA-seq. We are in process of evaluating divergence in early embryonic transcriptomes among sequenced strains in Drosophila melanogaster to assess the within species variation in gene expression. By contrasting within and between species divergence we will investigate the modes of evolution acting on gene expression during early fruifly embryogenesis. We will use our FlyFos system and quantitative imaging approaches to assess the divergence of spatial genes expression patterns before during and after the phylotypic period by imaging heterospecific transgenes from non-melanogaster species in the D. melanogaster environment. Our work will establish a new paradigm in post-genomic evolutionary developmental biology and reveal the evolutionary mechanism that underpins one of the most fundamental aspects of animal biology.
collaborators: Uwe Ohler (Duke University) and Casey Bergman (University of Manchester), Julia Jarrels (MPI-CBG Microarray facility)
funding: HFSP 2008, HFSP 2012 and ERC
Kalinka A. T., Varga K. M., Gerrard D. T., Preibisch S. W., Corcoran, D. L., Jarrels, J., Ohler, U., Bergman, C. M., Tomancak, P. Gene expression divergence recapitulates the developmental hourglass model Nature, 468, pp. 811-814, (2010)
Kalinka A.T., Tomancak P. (2012) The evolution of early animal embryos: conservation or divergence? Trends in Ecology and Evolution 27(7), 385-393