The VGP project at the MPI-CBG
The MPI-CBG and the CSBD are contributing to the international Vertebrate Genomes Project (VGP).
The VGP aims to generate error-free, near gapless reference-quality genome assemblies of all 66.000 vertebrate species. Obtaining the DNA sequences of all vertebrates, will enable the study of how genetic elements such as genes and regulatory regions have contributed to the evolution and fitness of these species.
The high-quality VGP genomes will become the main references for their species and will be stored in the Genome Ark, a digital open-access library. These genomes will be used to address novel questions, ranging from cell-type evolution to the genetics of complex traits and associated diseases. The Genome Ark will also provide tools for designing conservation strategies towards the preservation of life forms for future generations. Broadly, we expect that the VGP will provide a powerful resource to advance questions in biology, genomics, conservation, medicine, and bioinformatics.
The VGP at the MPI-CBG and the CSBD
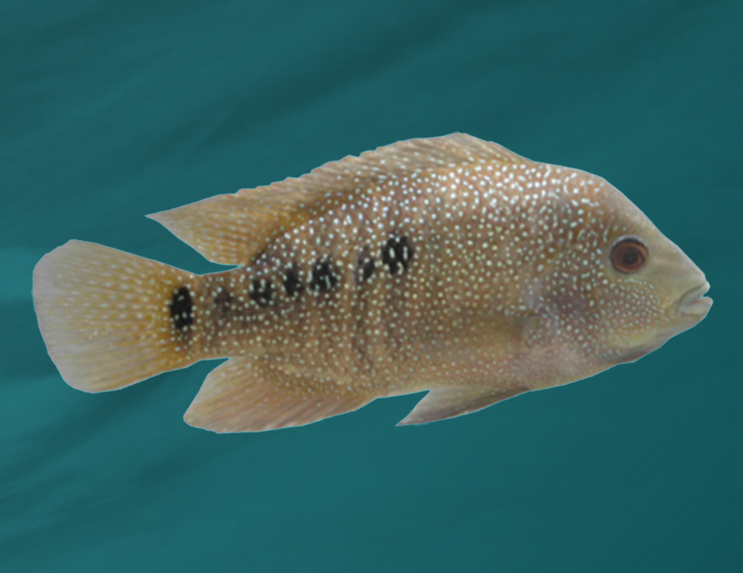
The MPI-CBG and the CSBD form one of the three international VGP sequencing hubs, together with the Rockefeller University, USA, and the Wellcome Sanger Genome Institute, UK. VGP in Dresden covers the sequencing, genome assembly and subsequent analysis of one representative species of each of the 260 vertebrate orders with a focus on bats and fish.
Apart from vertebrates for the VGP and Bat1K projects, we sequenced more vertebrates, invertebrates and plant species making use of long read technologies such as
a) other vertebrate species:
- very large vertebrate genomes of amphibians such as the Axolotl (Ambystoma mexicanum) and Spanish ribbed newt (Pleurodeles waltl),
- reptiles such as the tegu (Salvator merianae),
- fish such as the sand gopy (Pomatoschistus minutus) or a zebrafish cell line.
b) invertebrate species:
- five planarian species with highly AT-rich and repetitive genomes (Schmidtea mediterranea, Schmidtea polychroa, Polycelis tenius, Polycelis felina, Polycelis nigra),
- insects such as the cabbage fly (Delia radicum) and the hawk-moth (Hyles vespertilio) or
c) plants:
such as the wild tobacco (Nicotiana attenuata).
Species to be sequenced are selected between collaborators and tissues are submitted to the MPI-CBG, the CSBD, and the Dresden-concept Genome Center (DCGC).
A variety of de-novo sequencing technologies are currently applied and data are combined for genome assemblies to achieve our goal of error-free, near-gapless, chromosome-level, phased and annotated assemblies.
The current genome sequencing regime involves:
- 60x genome coverage of PacBio SMRT (single molecule real time DNA sequencing) reads,
- 68x genome coverage of 10x Genomics-linked reads for intermediate-range scaffolding,
- One DLS map making use of Bionano optical mapping to correct potential scaffolding errors,
- 68x HiC-linked reads for large-scale scaffolding,
- HiC and 10x Genomics linked short Illumina reads will be used for error correction of individual bases in the pipeline,
- RNAseq data and / or PacBio IsoSeq data for genome annotation.
PacBio SMRT DNA sequencing and 10x Genomics read cloud generation are done at the DCGC.
Bionano optical mapping will be established in Q4/2018 at the DCGC.
HiC is done exclusively with a commercial supplier (Arima Genomics, Inc. US).
The Genome assembly pipeline and Dazzler:
The Dresden genome assembling pipeline consists of two activities:
a) We are setting up a pipeline to generate error-free, near-gapless, chromosome-level and phased and assemblies making use of existing algorithms and software tools such as FALCON unzip, MARVEL, Scaff10X, TGH, Salsa, and, Arrow.
b) We are working on concepts and algorithms to analyze, understand and error correct long PacBio sequencing reads (The Dresden AZZembLER for long read DNA projects: dazzler). These pipelines will lead to a significant improvement of the assembly process in terms of accuracy, assembly continuity and finally required computing time.
dazzlerblog.wordpress.com